The use of single-cell multimodal omics technologies allows the profiling of multiple modalities from the same cell, providing a comprehensive view of cell states. The data also provide critical information about gene regulatory mechanisms. Much effort has been put into the development of single-cell multimodal omics methods. Current methods are limited by the complex workflow and low quality of data, which prevents the dissemination of the technology.
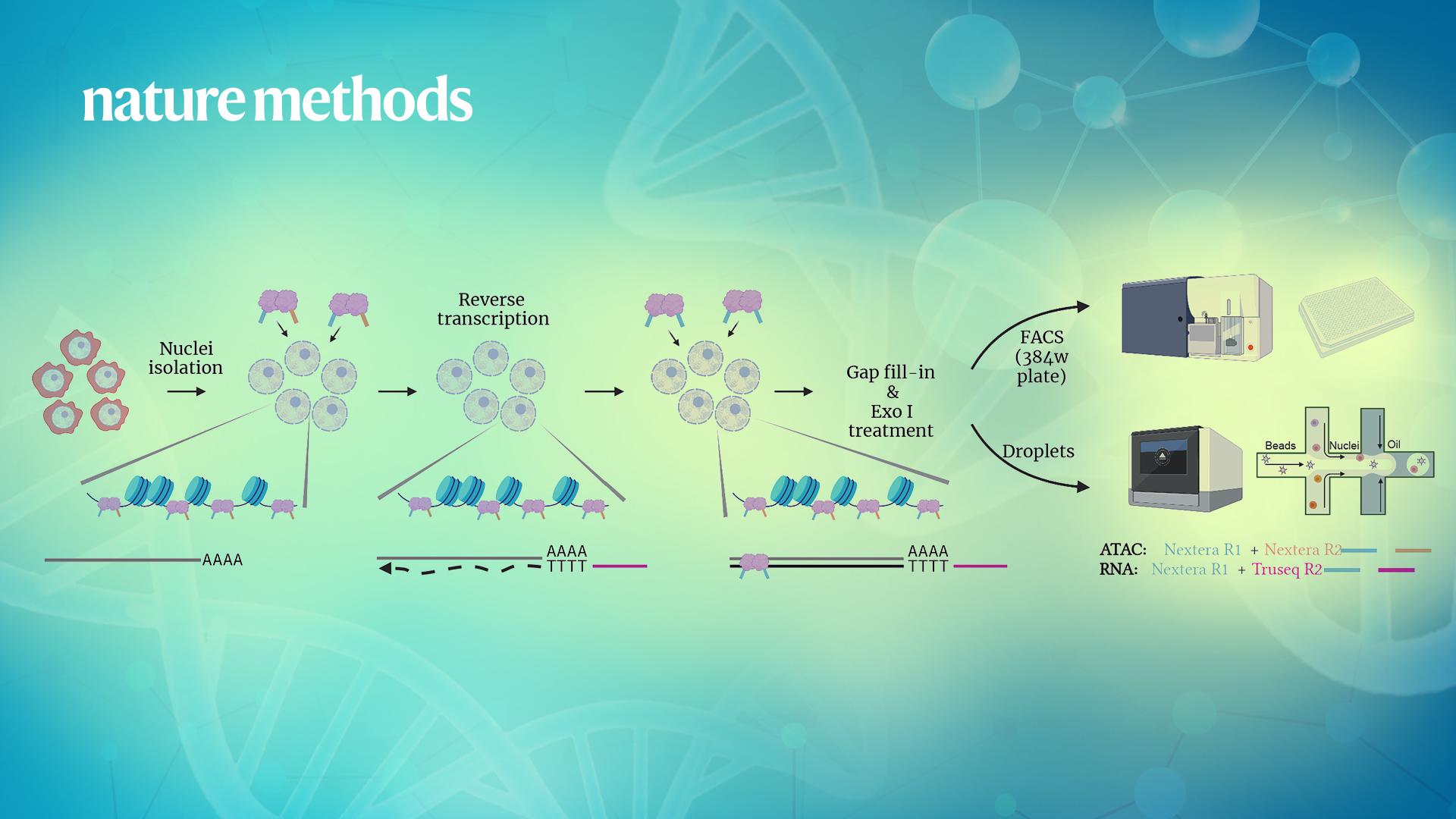
Researchers from Assistant Professor Xi Chen’s and Associate Professor Wenfei Jin’s labs in the School of Life Sciences at the Southern University of Science and Technology (SUSTech) have recently developed a sensitive, flexible, and modular single multimodal omics method called ISSAAC-seq. Their method enabled efficient multimodal profiling of chromatin accessibility and gene expression from the same cell. It can be used not only in the plate format for low throughput testing, but can also combine with droplet-based microfluidics for high throughput studies.
Their study, entitled “ISSAAC-seq enables sensitive and flexible multimodal profiling of chromatin accessibility and gene expression in single cells,” has been published in Nature Methods, a journal covering new basic research techniques in the life sciences.
ISSAAC-seq stands for in situ SHERRY after ATAC-seq. It took the advantage of a previously discovered property of the transpose Tn5 that it can cut RNA:DNA hybrids. Combined with the previously established single-cell ATAC-seq method, the ISSAAC-seq achieves the goal of simultaneous profiling of both chromatin accessibility and gene expression. The workflow includes: nuclei isolation -> open chromatin tagging -> in situ reverse transcription -> DNA:RNA hybrid tagging -> gap fill-in -> single cell isolation -> library preparation. This method used the differential barcoding strategy to mark open chromatin with the Nextera sequence and mRNA with the TruSeq sequence.
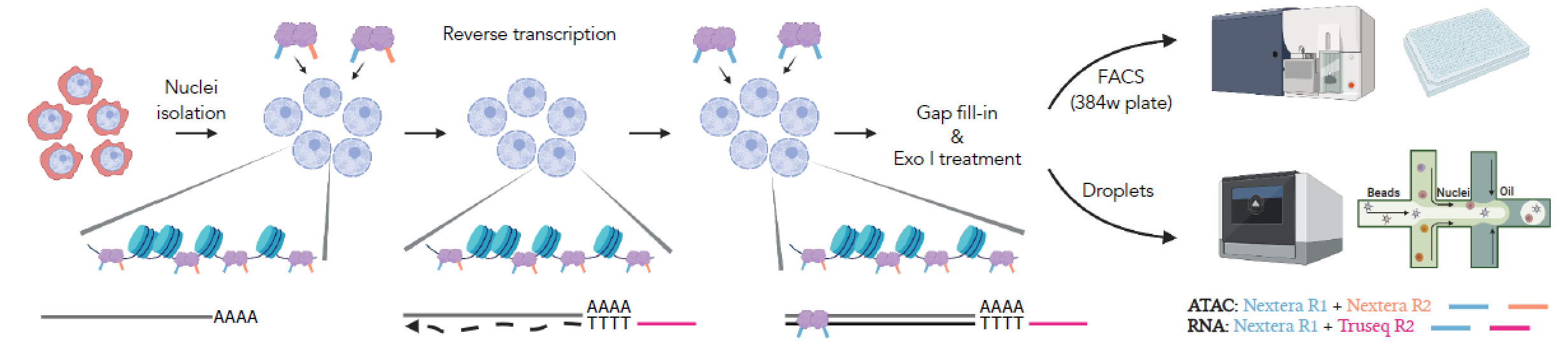
Figure 1. Schematic overview of the workflow of ISSAAC-seq
Such a workflow removed the necessity of multiple rounds of split-and-pool procedures in other methods, simplified the entire workflow and generated high-quality data. A big advantage of ISSAAC-seq is its modular design. ISSAAC-seq consists of two semi-independent modules: an in situ reaction module and a single-cell isolation module. The user can optimise based on their own need in each of the module separately, making the experiment very flexible. The researchers demonstrated that the data quality of ISSAAC-seq is much higher than previous open-source methods and comparable with the commercial platform. Still, the cost is only half or less than the commercial platform.
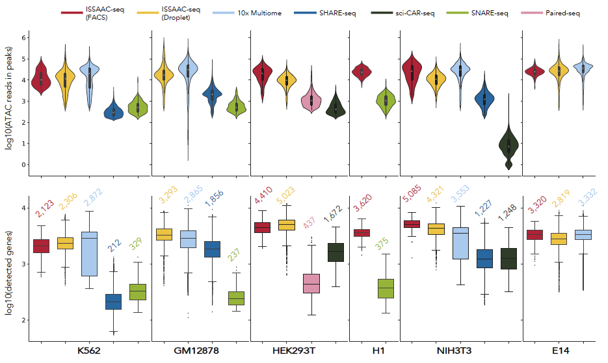
Figure 2. Data quality of ISSAAC-seq compared to other similar methods
Using ISSAAC-seq, the researchers analysed many mouse cortical neurons and revealed common and rare cell types in the mouse cortex together with their cell-type specific regulatory elements. They also uncovered the dynamics and relationship between chromatin accessibility and gene expression during an oligodendrocyte maturation trajectory.
In summary, ISSAAC-seq is robust and reproducible. The modular design makes it relatively straightforward for others to use and extend.
All of the authors from this study are from the School of Life Sciences at SUSTech. Research Asst. Prof. Wei Xu and Senior Research Scholar Weilong Yang are the co-first authors of this paper. Assoc. Prof. Wenfei Jin and Asst. Prof. Xi Chen are the co-corresponding authors.
The research was supported by the National Key R&D Program of China, National Natural Science Foundation of China (NSFC), Guangdong Basic and Applied Basic Research Foundation, Shenzhen Innovation Committee of Science and Technology, Shenzhen Science and Technology Program, and Shenzhen-Hong Kong Institute of Brain Science-Shenzhen Fundamental Research Institutions.
Paper link: https://www.nature.com/articles/s41592-022-01601-4
To read all stories about SUSTech science, subscribe to the monthly SUSTech Newsletter.
Proofread ByAdrian Cremin, Yingying XIA
Photo By