The transcription of genetic information from DNA to RNA by RNA polymerase (Pol) is a key step of the central dogma. In eukaryotes, besides the three conserved Pol I, Pol II, and Pol III, there are two specialized Pols, Pol IV and Pol V, in plants to produce non-coding RNA in the plant-specific RNA-directed DNA methylation pathway (RdDM).
While Pol IV functions in the biogenesis of pre-siRNA together with RDR2, Pol V functions in the biogenesis of long non-coding RNA (lncRNA) to serve as a scaffold for recruiting downstream gene silencing factors. Therefore, unlike other Pols, to produce and release the RNA transcripts, Pol V acts explicitly as a chromatin locator for other factors. The balance between the dynamic transcription elongation and stable chromatin binding of Pol V becomes a mystery.
A team led by Professor Jiamu Du of the Southern University of Science and Technology (SUSTech) has recently resolved this issue and published their results in the world-leading academic journal Science, entitled “Structure and mechanism of the plant RNA polymerase V”.
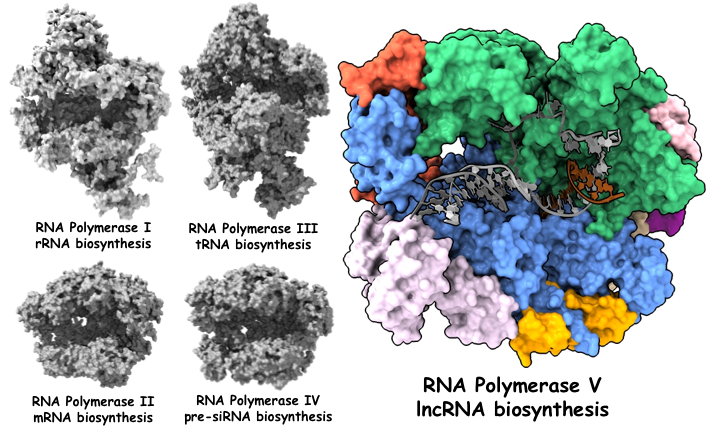
Figure 1. The five RNA polymerases in eukaryotes
The endogenous concentration of Pol V is extremely low, making it hard to be studied. According to the suggestion of Prof. Xiaofeng Cao from the Chinese Academy of Sciences (CAS), the research group used cauliflower as material to purify Pol V and determined its high-resolution cryo-EM structure. In the structure, Pol V uses a tyrosine residue to specifically stack with the dsDNA branch and to attenuate transcription elongation. The in vitro biochemical assay confirmed the blockage effects of transcription by Pol V. Moreover, the NPRE2 subunit of Pol V specifically captures the non-template strand DNA of the transcription bubble to enhance backtracking, yielding high 3’-5’ cleavage.
Overall, Pol V can both stall the transcription elongation and promote backtracking to make a balance between forward and backward moving of the transcription, preventing the transcription termination and ensuring the chromatin binding of Pol V.
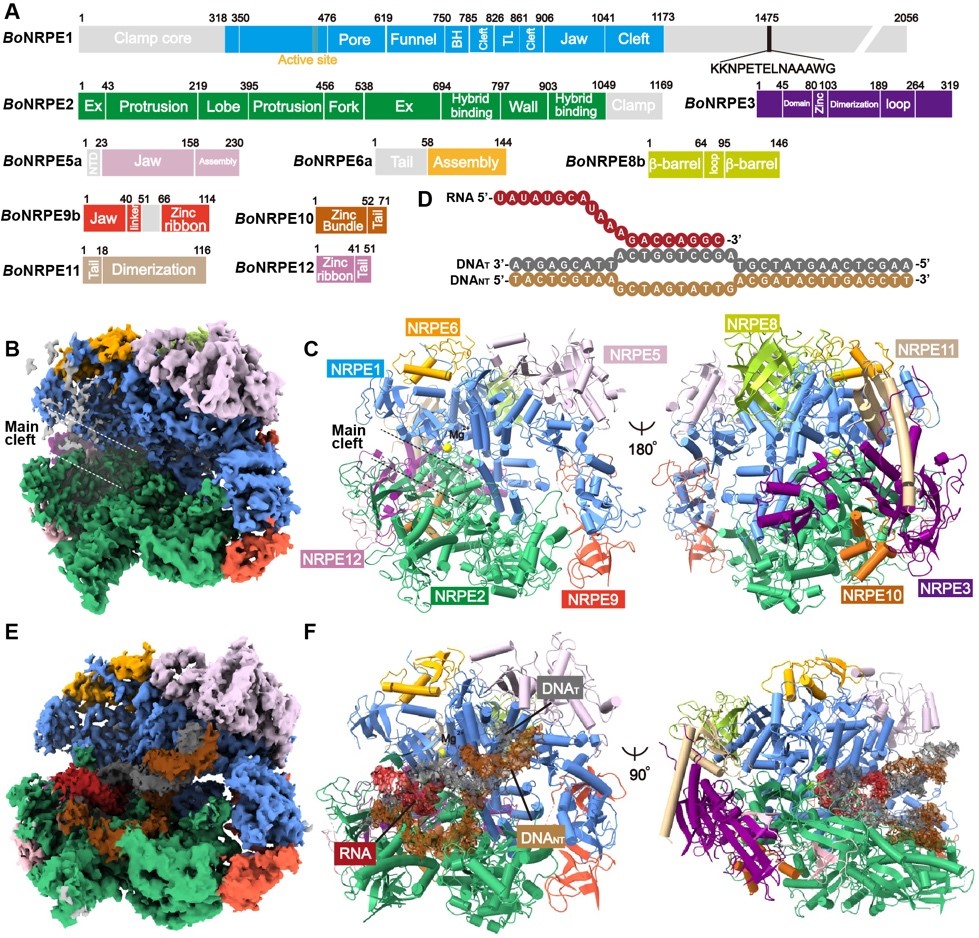
Figure 2. The overall structure of Pol V
As stated by a reviewer during revision, this work “finally completes the structural analysis of all 5 eukaryotic polymerases”, and provides direct evidence of how plant evolves its novel RNA polymerase adapted to the lncRNA production for efficient DNA methylation.
Mr. Guohui Xie, a Ph.D. student at SUSTech, Dr. Xuan Du, a visiting scholar from Shenzhen University (SZU), and Dr. Hongmiao Hu, a former visiting Ph.D. student at SUSTech, are the co-first authors of this paper. Prof. Jiamu Du is the corresponding author.
The Cryo-EM data was collected at the SUSTech Cryo-Electron Microscopy Center. Prof. Steven E. Jacobsen from the University of California, Los Angeles (UCLA), Prof. Xiaofeng Cao from CAS, and Prof. Sisi Li from SZU assisted with data processing and insightful discussions.
This research was supported by the Shenzhen Science and Technology Programs, National Natural Science Foundation of China, CAS, and NIH.
Paper link: https://www.science.org/doi/10.1126/science.adf8231
To read all stories about SUSTech science, subscribe to the monthly SUSTech Newsletter.
Proofread ByAdrian Cremin, Yingying XIA
Photo By