Recently, a research group led by Associate Professor Andrew P. Hutchins from the Department of Systems Biology, School of Life Sciences at the Southern University of Science and Technology (SUSTech) have published a paper titled “Dissecting the contribution of transposable elements to interphase chromosome structure” in journal Genome Biology. They developed a transposable element (TE)-focused Hi-C analysis pipeline, “te_hic,” and discovered for the first time that TEs are key determinants of 3D genome structure in the mammalian cell nucleus.
TE sequences make up nearly half of the human genome. They are capable of copying their own sequences and inserting them into other regions of the host genome, thereby achieving massive expansion during evolution. While most TEs have lost their transposition activity, recent studies indicate that they play important roles in biological processes such as gene regulation, embryonic development, and diseases. Meanwhile, the human genomic DNA is not linearly distributed in the cell nucleus, but rather folds into hierarchical and complex 3D chromatin structures, including A/B compartments, topologically associating domains (TADs), chromatin loops, and finer-scale promoter-enhancer contacts. These structures are closely related to transcriptional regulation and genomic stability.
Previous studies have suggested that certain TEs provide binding sites for key factors, such as CTCF, thus influencing chromatin loops and TAD boundaries. However, there is a lack of systematic answers regarding the genome-wide contribution of TEs to 3D chromatin interactions and the underlying mechanisms. A core obstacle is that traditional Hi-C analysis pipelines generally discard or de-emphasize multi-aligned reads on repetitive sequences, resulting in a long-standing failure to fully incorporate TE-related chromatin contacts into systematic analyses.
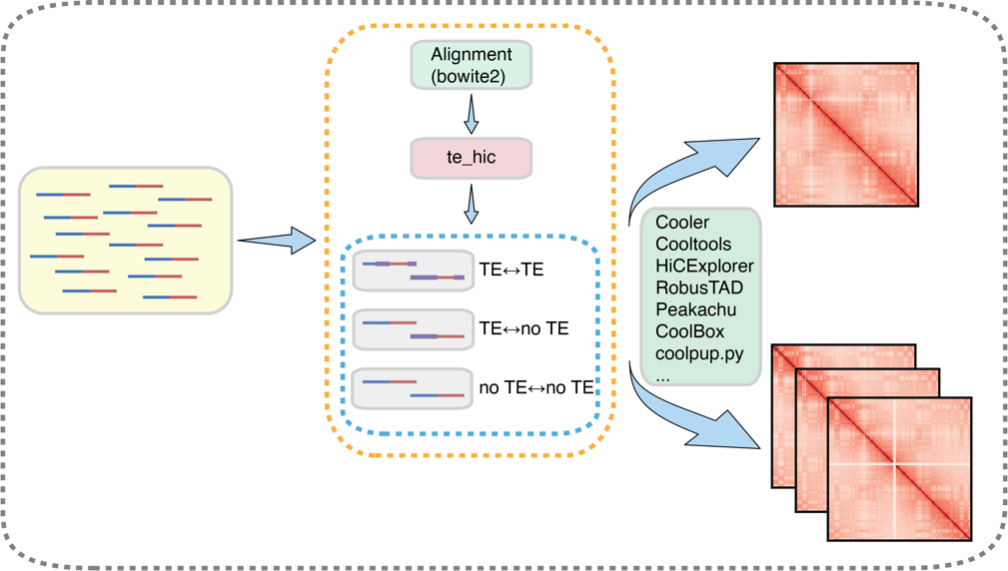
Figure 1. “te_hic” pipeline
The study revealed that using only TE-containing Hi-C reads can still substantially reconstruct the main features of the genome-wide Hi-C matrix. Notably, classic 3D genome structures, such as A/B compartments and TADs, can be reconstructed solely through reads containing TE sequences, suggesting that TEs are an important source of 3D chromatin structural signals. At a quantitative level, the study further estimated that when considering only TE-TE interactions, TEs can account for approximately 30% of the 3D genome structure; their contribution reaches approximately 78% if TE-no TE interactions are included. These results demonstrate that TE sequences occupy a critical position in chromatin folding within the interphase nucleus.
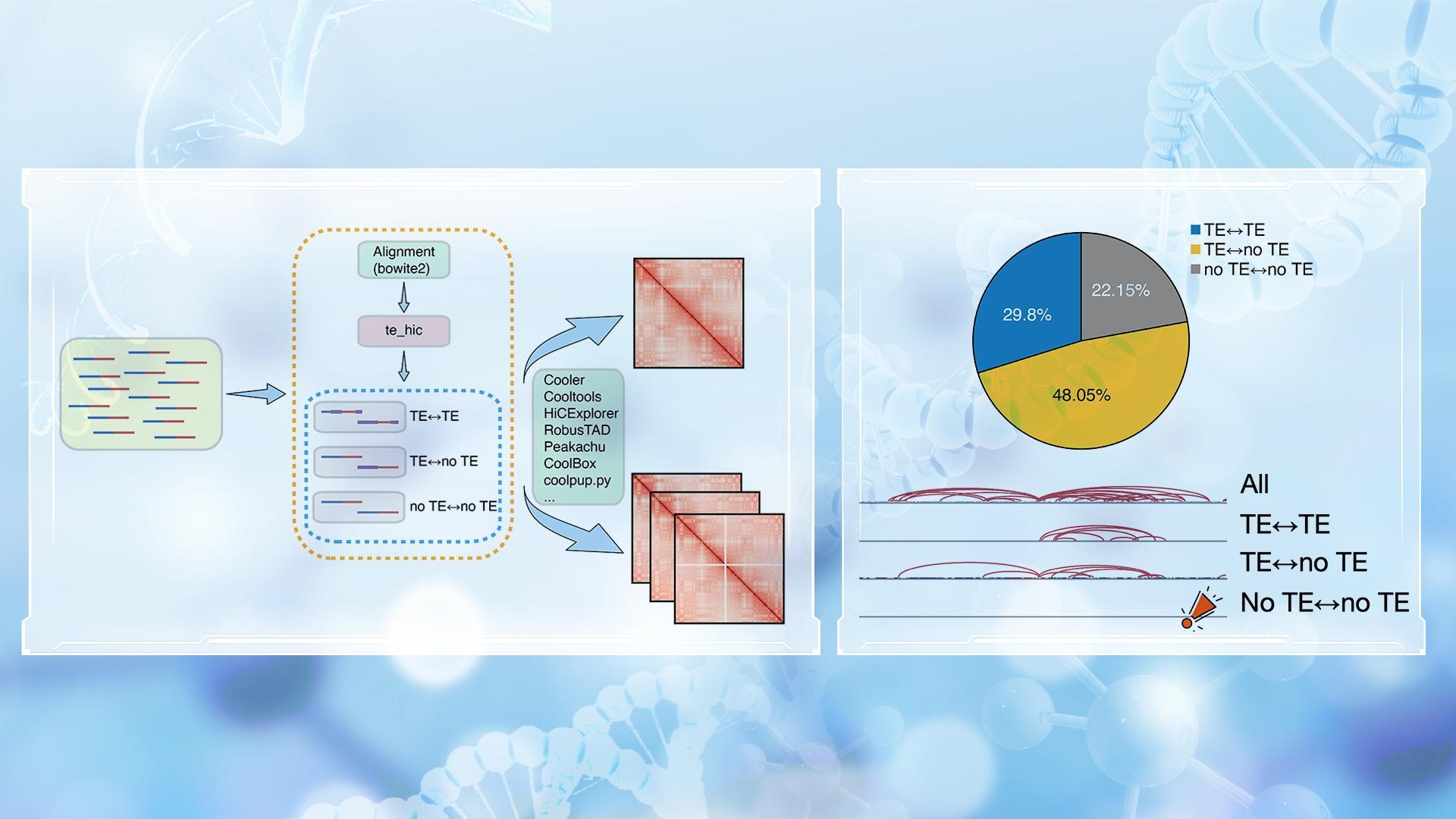
To systematically evaluate the role of transposable elements in the 3D genome, Dr. Hutchins’s team developed te_hic, which incorporates TE-related Hi-C reads into downstream analysis. Unlike conventional Hi-C pipelines, te_hic specifically retains and integrates TE-related reads during the alignment and Hi-C matrix construction stages. This allows the pipeline to directly focus on “TE-TE” and “TE-no TE” chromatin interactions, thereby more precisely reflecting the contribution of TE sequences within the 3D genome.
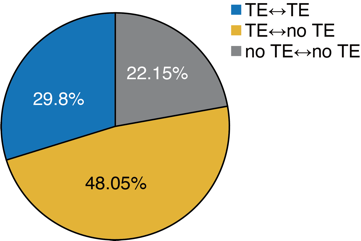
Figure 2. Proportion of Hi-C read pairs with TEs at their ends
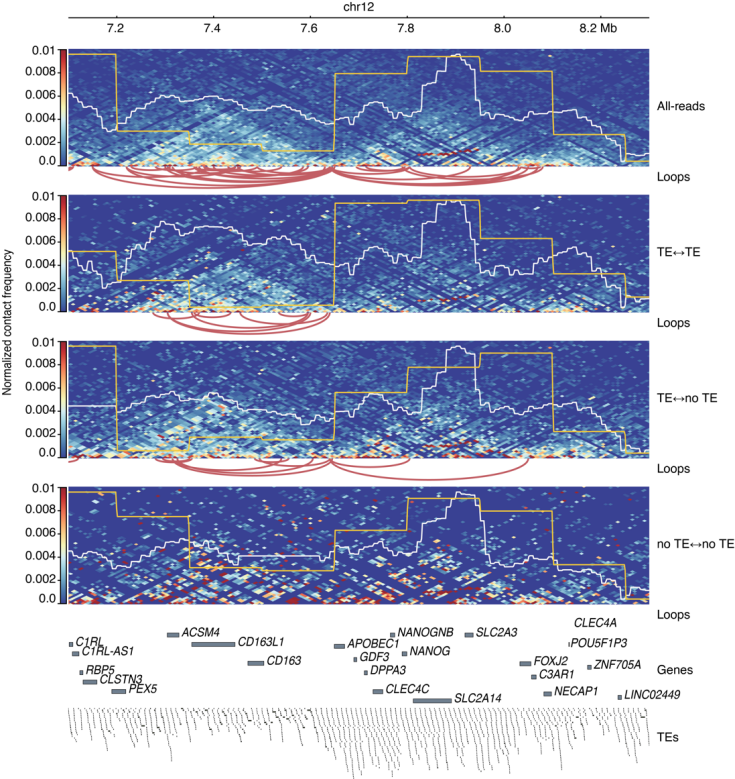
Figure 3. Triangle contact heatmaps showing TEs as are key determinants of 3D genome structure
The research team subsequently integrated multiple DNA-binding factor data and Hi-C data from human and mouse pluripotent stem cells (PSCs), proposing that TEs shape 3D genome structure by recruiting epigenetic or transcription factors (ETFs). Further computational analysis identified a class of classic factors positively correlated with chromatin interactions (contact-positive), such as CTCF, RAD21, and SMC3. Meanwhile, they also found that certain factors are associated with decreased local chromatin interactions (contact-negative), such as RNF2. Furthermore, they identified new contact-positive regulatory factors, such as SMARCA4 and MAFK, revealing that TE-mediated 3D interactions are likely driven by multiple classes of factors rather than being strictly limited to a CTCF-dependent mechanism.
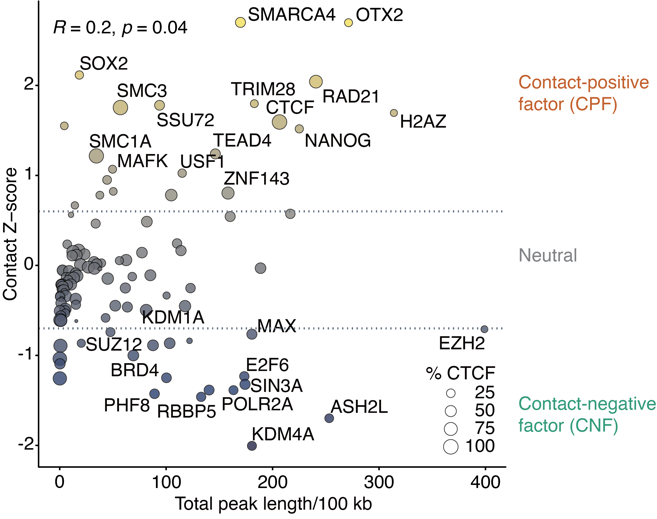
Figure 4. Transcription factors and Epi-factors can be classified in to three groups based on their association with chromatin contacts.
To validate these computational predictions, the authors performed knockdown (KD) experiments on candidate factors followed by Hi-C assays. The results showed that knocking down SMARCA4 or MAFK led to a reduction in TE-mediated chromatin interactions, providing experimental evidence for their positive regulation of TE-related 3D chromatin structures. Notably, the knockdown of SMARCA4 had a more significant impact on short-range contacts, suggesting that it may maintain local 3D structural stability by binding to TEs.
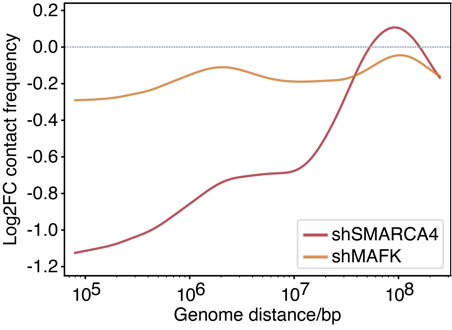
Figure 5. Knockdown of contact-positive factors reduce short-range chromatin contacts.
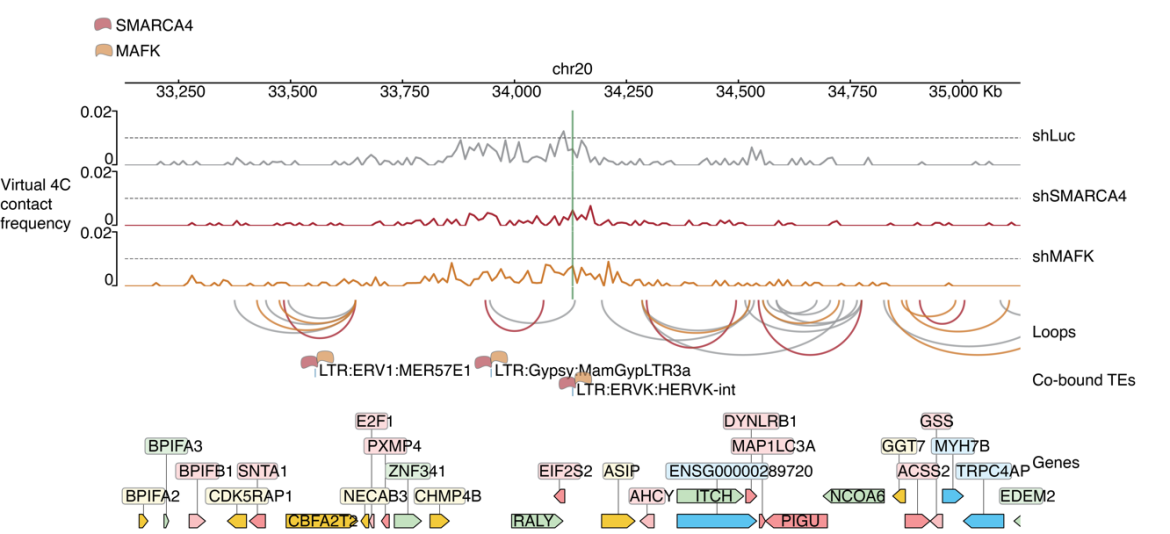
Figure 6. Decreased chromatin contacts at a TE locus bound by both SMARCA4 and MAFK after their knockdown.
Overall, this study advances the understanding of the 3D genome from both methodological and mechanistic perspectives. On one hand, te_hic provides a reusable analysis tool for studying the involvement of repetitive sequences in the 3D genome, reducing the systematic loss of TE signals in traditional analysis pipelines. On the other hand, the research proposes and validates a broader mechanistic framework of “TE–ETF binding–chromatin interaction,” indicating that TEs are key determinants of 3D genome structure in mammalian cells. These findings provide new insights for understanding 3D chromatin structure and its associated gene expression and epigenetic regulation, as well as laying a foundation for further dissecting the regulatory functions and molecular mechanisms of TEs across multiple biological processes.
SUSTech Master’s student Liyang SHI (currently a PhD candidate in Kyoto-McGill International Joint Ph.D. Program in Genomic Medicine), postdoctoral fellow Zhen XIAO, and Research Assistant Professor Xuemeng ZHOU are co-first authors of the paper. Associate Professor Andrew P. Hutchins is the corresponding author, with SUSTech as the primary affiliated institution.
Proofread ByNoah Crockett, Junxi KE
Photo ByYan QIU