Quanying LIU’s team from the Department of Biomedical Engineering at Southern University of Science and Technology (SUSTech), under the guidance of Academician Dongfeng GU at SUSTech, has made new progress in the field of brain source imaging. The related findings, titled “A geometry aware framework enhances noninvasive mapping of whole human brain dynamics,” have been published in Nature Biomedical Engineering. The study addresses the long-standing ill-posed inverse problem and the lack of biologically grounded priors in EEG/MEG source imaging, and proposes a geometry-aware source imaging framework, Geometric Basis Function (GBF).

EEG and MEG can record brain activity noninvasively with millisecond temporal resolution. However, scalp potentials and extracranial magnetic fields are mixed observations generated by activity from multiple brain regions after volume conduction through the head, and therefore cannot directly reveal the location or strength of cortical neural sources. EEG/MEG source imaging aims to infer brain activity from sensor-level signals, but this inverse problem is highly ill-posed. Conventional methods often rely on generic mathematical priors such as minimum energy or spatial smoothness, which do not fully capture individual anatomical differences.
The core idea of GBF is to incorporate individual brain structure directly into source imaging. The method reconstructs each participant’s cortical surface from structural MRI and extracts Laplace-Beltrami eigenmodes that capture individual cortical geometry. These modes serve as geometric basis functions, allowing neural activity at any time point to be represented as a linear combination of a small number of individualized basis functions. As a result, source reconstruction is constrained within a space consistent with each participant’s cortical geometry, improving stability, spatial precision, and anatomical interpretability.
Within the GBF framework, the study first reconstructs individual cortical surfaces from structural MRI and derives geometric eigenmodes through Laplace-Beltrami decomposition. These modes describe spatial patterns of cortical activity and constrain EEG/MEG source imaging within the geometry defined by the individual brain (Fig. 1a-c). The team then validated GBF at multiple levels; source localization was evaluated on the Meta-Source Benchmark (Fig. 1d), task-evoked EEG data were used to test whether GBF could recover canonical sensory and motor cortical activations (Fig. 1e), and further applications included MEG-derived virtual iEEG connectome reconstruction, intracranial stimulation propagation analysis, and localization of epilepsy-related brain regions (Fig. 1f-h).

Figure 1. EEG/MEG source reconstruction framework constrained by geometric basis functions
Because the true neural source distribution in real EEG data is usually unknown, source imaging methods are difficult to validate directly. Traditional simulation datasets often rely on random point sources or localized activation patches, which cannot fully reflect the complex spatial patterns of real brain activity. To address this limitation, the study constructed the Meta-Source Benchmark, a biologically interpretable dataset with known reference source distributions for systematic evaluation of EEG/MEG source imaging methods.
The dataset was built from 26,273 fMRI statistical images from NeuroVault and 1,307 cognitive terms from Neurosynth. Dimensionality reduction and clustering were used to generate 200 representative cortical source maps, which were then projected through head models to produce corresponding EEG scalp topographies. This resulted in 200 source-EEG pairs with known source distributions. Compared with conventional simulations, the Meta-Source Benchmark better approximates realistic spatial patterns of brain activity and provides functionally annotated ground truth for source localization evaluation.
Using this benchmark, the study compared GBF with several conventional source imaging methods under both Gaussian noise and realistic EEG noise. GBF achieved better performance across NRMSE, localization error, Pearson correlation, cosine similarity, and AUC, demonstrating higher source localization accuracy and stronger noise robustness (Fig. 2b-d).

Figure 2. Construction and evaluation of the Meta-Source Benchmark
After benchmark validation, the study applied GBF to real task-evoked EEG data to further test its generalizability across visual, auditory, somatosensory, and motor tasks (Fig. 3a). Sensor-level analyses showed clear responses consistent with established neurophysiology. Visual, auditory, and somatosensory tasks produced distinct ERP/GFP peaks, while the motor tasks showed ERD/ERS changes related to movement execution (Fig. 3b).
Using task-related Neurosynth meta-analytic maps as spatial references, GBF produced source reconstructions that showed stronger correspondence with expected task-related regions. The visual, auditory, somatosensory, and motor tasks were localized to the primary visual cortex, bilateral auditory cortices, contralateral somatosensory cortex, and left motor cortex, respectively (Fig. 3c).
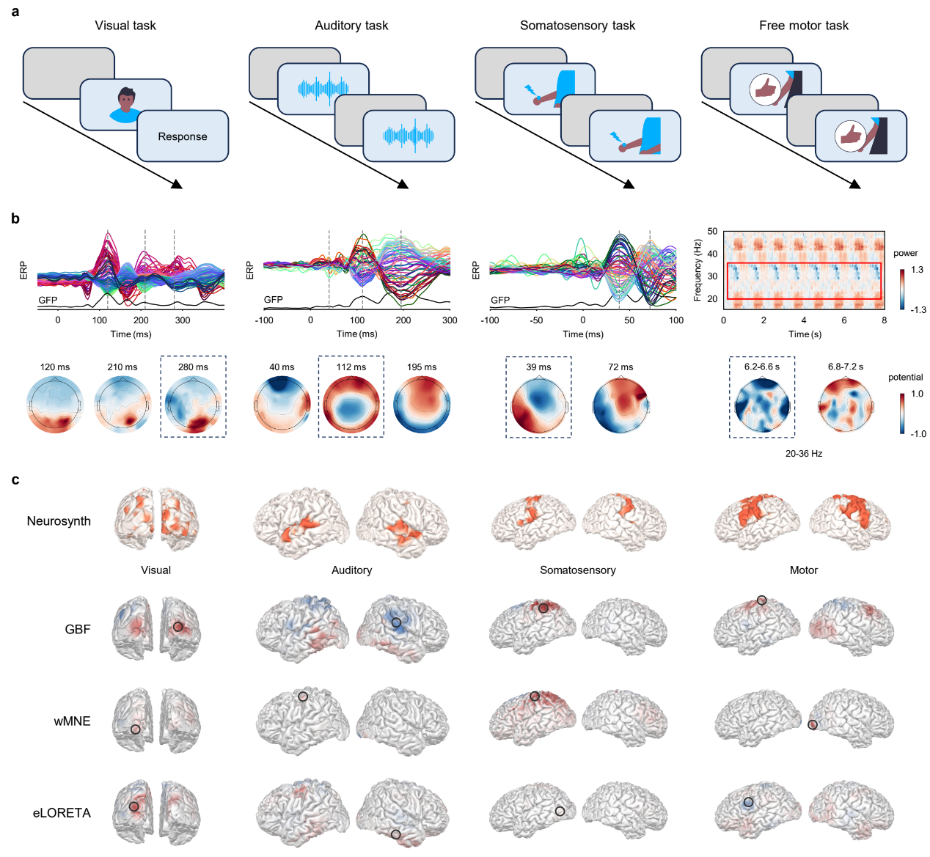
Figure 3. GBF captures task-evoked neural activity patterns
Resting-state functional connectivity reflects whole-brain network organization. To assess whether GBF can recover source-space connectivity from noninvasive MEG signals, the study compared MEG-derived virtual intracranial EEG (ViEEG) connectomes with real intracranial EEG (iEEG) connectomes (Fig. 4a).
Specifically, the study constructed a group-level iEEG connectivity matrix from resting-state iEEG data of 110 presurgical epilepsy patients, and reconstructed ViEEG connectomes from resting-state MEG data of 80 healthy HCP participants using different source imaging methods. GBF achieved the highest iEEG-ViEEG correlations in the theta, alpha, beta, and low-gamma bands, and matched MNE in the delta band (Fig. 4b).
Spatial null-model analysis further showed that the connectivity patterns recovered by GBF were significantly above random distributions, indicating genuine electrophysiological agreement between MEG and iEEG. GBF performed particularly well in the beta band and reproduced the spatial distribution of high-strength iEEG edges (Fig. 4c-e).
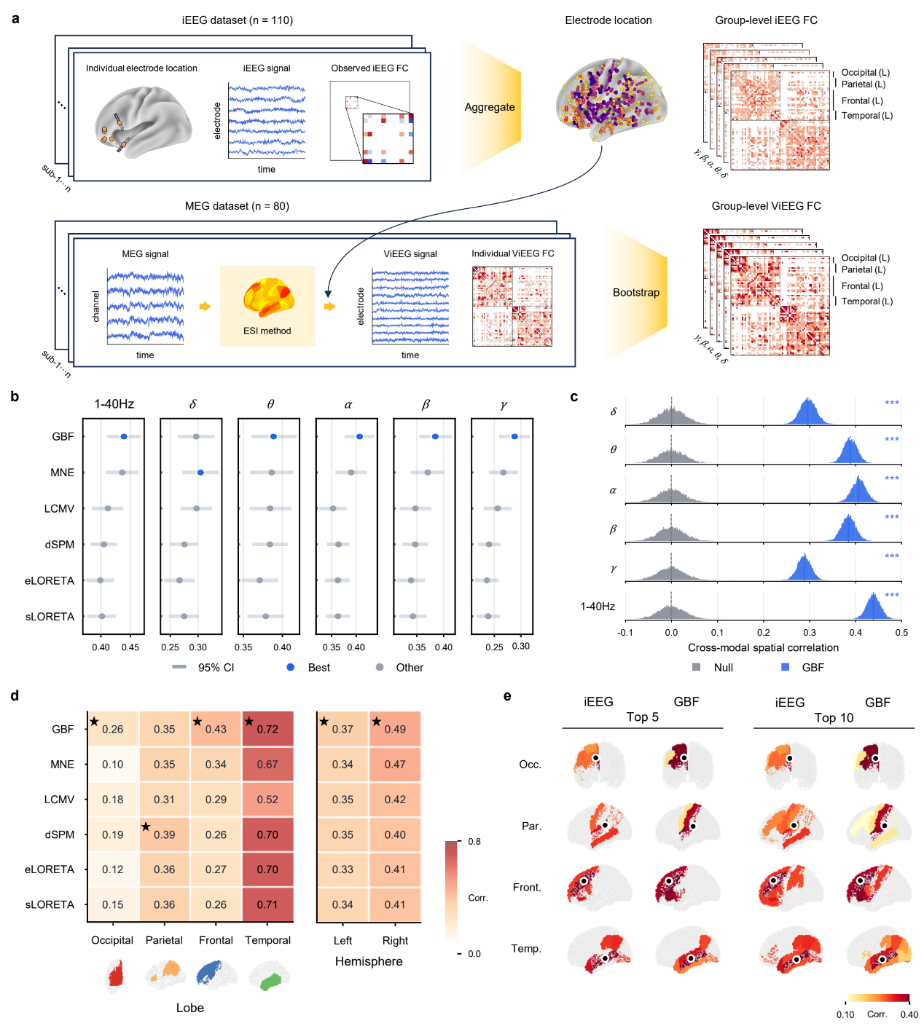
Figure 4. GBF reconstructs resting-state electrophysiological functional connectomes
To validate GBF in a real perturbation setting, the study applied the method to simultaneous intracranial electrical stimulation (iES) and high-density scalp EEG data. Because the stimulation sites are known, this dataset provides an approximate reference for assessing whether source imaging methods can reconstruct stimulation-evoked cortical responses from scalp EEG (Fig. 5a).
During the experiment, intracranial electrodes delivered electrical pulses to specific brain regions, inducing N1 and N2 responses in scalp EEG. The study compared the spatial distance between the reconstructed activation peaks and the true stimulation sites. Across 35 participants and 318 stimulation sessions, GBF showed lower localization error than conventional methods and remained robust across different stimulation depths (Fig. 5b-e).
Beyond stimulation-site localization, the study used GBF to reconstruct cortical propagation after stimulation. By combining phase-gradient and optical-flow analyses, GBF captured rapid directed propagation from the stimulation site during the N1 window and broader, slower diffusion during the N2 window (Fig. 5f). These results indicate that GBF can not only localize stimulation-evoked activity, but also characterize whole-brain spatiotemporal propagation dynamics.

Figure 5. GBF localizes intracranial stimulation sites and tracks stimulation-evoked neural propagation
Localization of epilepsy-related brain regions is a key step in presurgical evaluation for drug-resistant epilepsy. To assess the clinical potential of GBF, the study first validated the method on a public dataset of 24 patients with favorable postsurgical outcomes, using the postoperative resection extent as an approximate reference for the epileptogenic zone (EZ). GBF was applied to reconstruct sources from interictal epileptiform discharges (Fig. 6a).
The results showed that GBF activation peaks were closer to the resection area and showed a consistent advantage over conventional methods (Fig. 6b). Representative cases demonstrated good spatial agreement between GBF-reconstructed cortical activation and postoperative resection masks (Fig. 6c,d).
The study further analyzed two 256-channel high-density EEG cases from Huashan Hospital, Fudan University, and two publicly available 19-channel low-density EEG cases. GBF early-source peaks were closer to clinically annotated seizure-onset zones (SOZs) and showed more focal spatial patterns (Fig. 6e,f). These findings suggest that GBF has clinical potential for noninvasive localization of epilepsy-related brain regions.

Figure 6. GBF facilitates noninvasive localization of epilepsy-related brain regions
Overall, EEG/MEG source imaging has long been limited by the ill-posed nature of the inverse problem, often leading to spatially blurred reconstructions and reduced biological interpretability. The GBF framework embeds individual cortical geometry directly into the source imaging process, using anatomical constraints to improve the spatial accuracy, stability, and interpretability of whole-brain neural dynamics reconstruction.
The key feature of GBF is the transformation of MRI-derived cortical surfaces into geometric basis functions through Laplace-Beltrami decomposition. Unlike conventional methods based on spherical assumptions, localized patch priors, or region-level connectome modes, GBF models neural sources directly on each individual’s native cortical surface, thereby capturing individual anatomical variability. Its closed-form MAP solution also makes the method computationally efficient and straightforward to implement.
Systematic validation showed that GBF achieved higher reconstruction accuracy and noise robustness on the Meta-Source Benchmark, recovered canonical sensory and motor areas in task-evoked EEG, better reproduced iEEG functional connectivity structures in resting-state MEG analysis, localized stimulation sites, and tracked millisecond-scale propagation in intracranial stimulation data, and demonstrated potential for assisting noninvasive epilepsy localization. Together, these findings indicate that individual cortical geometry improves source localization and provides an effective constraint for analyzing whole-brain network dynamics.
In the future, the GBF framework may be extended to subcortical structures such as the hippocampus and thalamus, and combined with temporal priors or state-space models to better characterize continuous neural propagation. Geometric basis functions may also serve as compact, individualized brain-structure representations for integration with deep learning models, reducing model complexity while improving biological plausibility.
The paper was co-first-authored by Song WANG, Master’s student at SUSTech; Kexin LOU, joint PhD candidate from SUSTech and the University of Queensland; Chen WEI, joint PhD candidate from SUSTech and the University of Birmingham. Liang CHEN, Chief Physician at Huashan Hospital, Fudan University; Dongfeng GU, Academician at SUSTech; and Quanying LIU, Associate Professor at SUSTech, are the co-corresponding authors. The related technology has been filed as an invention, and patent applications have been formally accepted. Co-founded by the three co-first authors, Song WANG, Kexin LOU, and Chen WEI from SUSTech, to promote applications in noninvasive brain functional imaging and clinical neurological assessment.
Paper Link: https://www.nature.com/articles/s41551-026-01664-0
Proofread ByNoah Crockett, Junxi KE
Photo By